Overview — basic information
| Identifier | S1 |
|---|---|
| Type | Enzyme family |
| Number of CGCs | 33,540 |
| Number of proteins (in associated CGCs) | 0 |
| Number of S1 proteins (target-only) | 0 |
| Source data files | cgcs.txt · sequences.faa · proteins.tsv |
Overview — data sources
| CGC catalog | all_cgc_magmapped.tsv · 2.45 M rows / 24,195 MAGs |
|---|---|
| All-vs-all DIAMOND | 873.6 M edges (e ≤ 1e-30, k 1000) |
| Annotation source | dbCAN (CAZyme), SulfAtlas (sulfatases), TC/TF/STP/Peptidase metadata |
| GTDB taxonomy | gomc_metadata.tsv · mag_taxonomy.tsv |
| NMPF cross-reference | gomc_nmpfs.tsv |
Phylogeny
FastTree (Newick) not yet built. Once msa_fasttree.py finishes, this section will host an interactive phylotree.js viewer.
Most common taxonomy
Genome counts hosting at least one S1 CGC, drilled down through GTDB ranks. Coloured by the 11-group palette used in the rarefaction and UMAP figures (grey = "other").
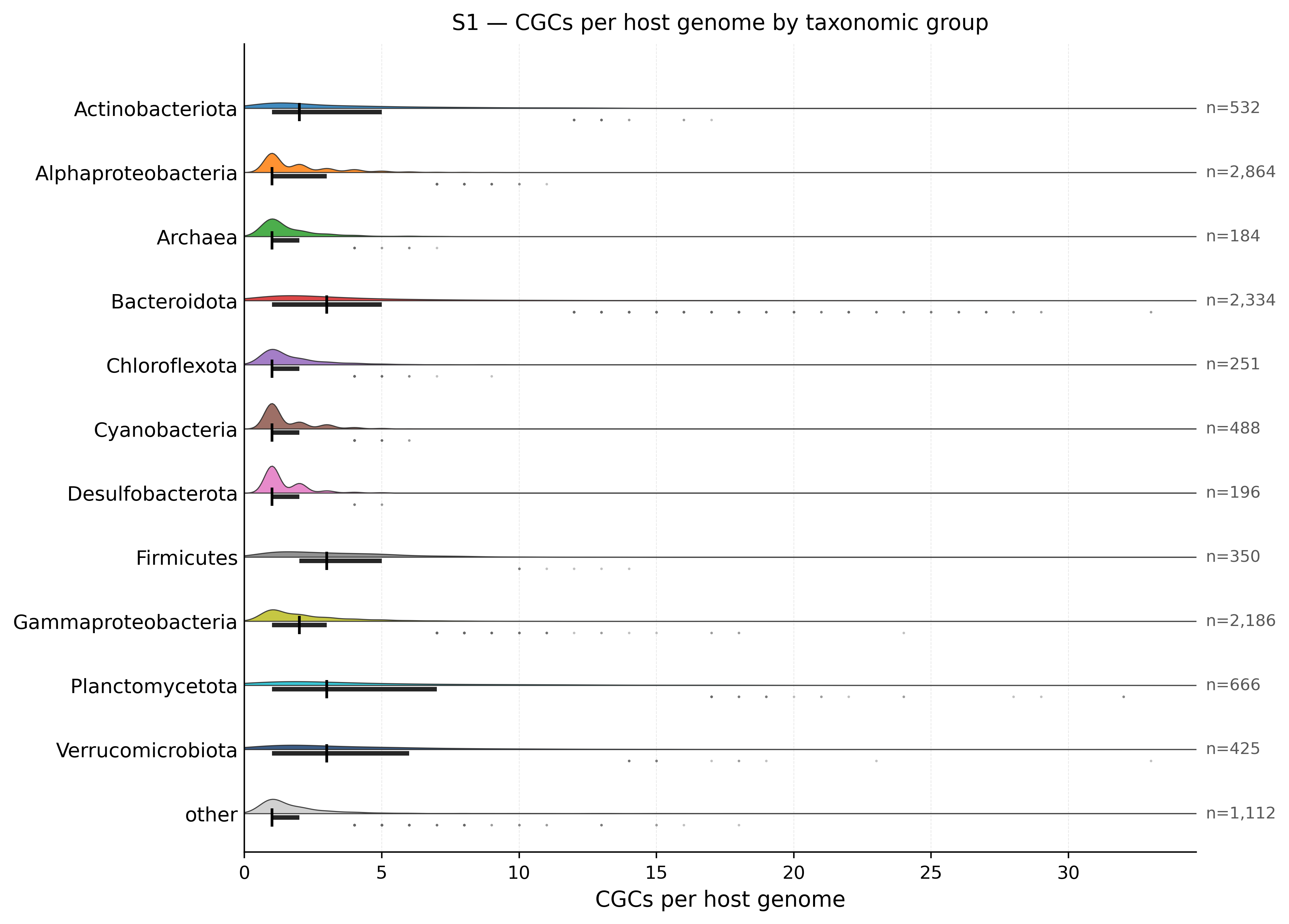
Host taxonomic groups — CGCs per genome
Half-violin density distribution of CGCs per host genome (restricted to S1-containing CGCs), split by the 11 reference taxonomic groups used in the rarefaction figure. Median + IQR shown as black tick & bar; outliers (1.5×IQR) as small dots.
Gene neighborhood — co-occurring families
First-family token per protein (GH / PL / CE / S1 only). Top 30 of 571.
| Family | Count |
|---|---|
| S1 | 41,181 |
| S1_6 | 4,401 |
| S1_8 | 3,497 |
| S1_7 | 3,090 |
| S1_2 | 3,066 |
| GH13 | 2,923 |
| GH3 | 2,333 |
| S1_4 | 2,244 |
| S1_16 | 2,232 |
| S1_15 | 2,184 |
| GH2 | 1,840 |
| S1_27 | 1,838 |
| GH43 | 1,715 |
| S1_11 | 1,697 |
| S1_14 | 1,516 |
| S1_37 | 1,388 |
| GH29 | 1,175 |
| GH16 | 1,100 |
| S1_17 | 1,032 |
| S1_9 | 1,030 |
| GH20 | 1,006 |
| GH92 | 933 |
| S1_22 | 866 |
| S1_97 | 846 |
| GH23 | 844 |
| S1_12 | 839 |
| S1_20 | 798 |
| S1_0 | 782 |
| GH5 | 760 |
| CE9 | 755 |
SCoNe — co-occurrence network
SCoNe artefacts not yet built.
Associated CGCs
Showing first 80 of 33,540 CGCs. Download full list.
| CGC ID | MAG |
|---|---|
| BATS_SAMN07137085_METAG_DFMHGLCG_CGC10 | BATS_SAMN07137085_METAG_DFMHGLCG |
| BATS_SAMN07137085_METAG_DFMHGLCG_CGC13 | BATS_SAMN07137085_METAG_DFMHGLCG |
| BATS_SAMN07137085_METAG_DFMHGLCG_CGC7 | BATS_SAMN07137085_METAG_DFMHGLCG |
| BATS_SAMN07137085_METAG_DFMHGLCG_CGC8 | BATS_SAMN07137085_METAG_DFMHGLCG |
| BATS_SAMN07137116_METAG_OMDFKJGI_CGC9 | BATS_SAMN07137116_METAG_OMDFKJGI |
| BATS_SAMN07137118_METAG_HPJDOJIB_CGC1 | BATS_SAMN07137118_METAG_HPJDOJIB |
| BATS_SAMN07137118_METAG_HPJDOJIB_CGC14 | BATS_SAMN07137118_METAG_HPJDOJIB |
| BATS_SAMN07137118_METAG_HPJDOJIB_CGC15 | BATS_SAMN07137118_METAG_HPJDOJIB |
| BATS_SAMN07137118_METAG_HPJDOJIB_CGC2 | BATS_SAMN07137118_METAG_HPJDOJIB |
| BATS_SAMN07137118_METAG_HPJDOJIB_CGC9 | BATS_SAMN07137118_METAG_HPJDOJIB |
| BATS_SAMN08390925_METAG_BJAOANMI_CGC11 | BATS_SAMN08390925_METAG_BJAOANMI |
| BATS_SAMN08390925_METAG_BJAOANMI_CGC18 | BATS_SAMN08390925_METAG_BJAOANMI |
| BATS_SAMN08390925_METAG_BJAOANMI_CGC4 | BATS_SAMN08390925_METAG_BJAOANMI |
| BATS_SAMN08390925_METAG_BJAOANMI_CGC9 | BATS_SAMN08390925_METAG_BJAOANMI |
| BGEO_SAMN07136507_METAG_NAJFPCKG_CGC13 | BGEO_SAMN07136507_METAG_NAJFPCKG |
| BGEO_SAMN07136507_METAG_NAJFPCKG_CGC6 | BGEO_SAMN07136507_METAG_NAJFPCKG |
| BGEO_SAMN07136507_METAG_NAJFPCKG_CGC7 | BGEO_SAMN07136507_METAG_NAJFPCKG |
| BGEO_SAMN07136507_METAG_NAJFPCKG_CGC8 | BGEO_SAMN07136507_METAG_NAJFPCKG |
| BGEO_SAMN07136523_METAG_APAOEKCM_CGC1 | BGEO_SAMN07136523_METAG_APAOEKCM |
| BGEO_SAMN07136523_METAG_NMOKDPNF_CGC7 | BGEO_SAMN07136523_METAG_NMOKDPNF |
| BGEO_SAMN07136546_METAG_CABEIBHM_CGC1 | BGEO_SAMN07136546_METAG_CABEIBHM |
| BGEO_SAMN07136583_METAG_OEAOIHAM_CGC13 | BGEO_SAMN07136583_METAG_OEAOIHAM |
| BGEO_SAMN07136583_METAG_OEAOIHAM_CGC15 | BGEO_SAMN07136583_METAG_OEAOIHAM |
| BGEO_SAMN07136583_METAG_OEAOIHAM_CGC16 | BGEO_SAMN07136583_METAG_OEAOIHAM |
| BGEO_SAMN07136583_METAG_OEAOIHAM_CGC2 | BGEO_SAMN07136583_METAG_OEAOIHAM |
| BGEO_SAMN07136681_METAG_IEGHCJCC_CGC6 | BGEO_SAMN07136681_METAG_IEGHCJCC |
| BGEO_SAMN07136689_METAG_KJMIPNPG_CGC2 | BGEO_SAMN07136689_METAG_KJMIPNPG |
| BGEO_SAMN07136708_METAG_NLOEKJNE_CGC2 | BGEO_SAMN07136708_METAG_NLOEKJNE |
| BGEO_SAMN07136709_METAG_EEKAFEOM_CGC2 | BGEO_SAMN07136709_METAG_EEKAFEOM |
| BGEO_SAMN07136710_METAG_IMMBNPOM_CGC10 | BGEO_SAMN07136710_METAG_IMMBNPOM |
| BGEO_SAMN07136710_METAG_IMMBNPOM_CGC11 | BGEO_SAMN07136710_METAG_IMMBNPOM |
| BGEO_SAMN07136788_METAG_AFBPEAAE_CGC21 | BGEO_SAMN07136788_METAG_AFBPEAAE |
| BGEO_SAMN07136788_METAG_AFBPEAAE_CGC24 | BGEO_SAMN07136788_METAG_AFBPEAAE |
| BGEO_SAMN07136898_METAG_IGLCDHNE_CGC1 | BGEO_SAMN07136898_METAG_IGLCDHNE |
| BGEO_SAMN07136920_METAG_MGPENMLJ_CGC6 | BGEO_SAMN07136920_METAG_MGPENMLJ |
| BGEO_SAMN07136920_METAG_MGPENMLJ_CGC7 | BGEO_SAMN07136920_METAG_MGPENMLJ |
| BGEO_SAMN07136937_METAG_ALPGGBFO_CGC13 | BGEO_SAMN07136937_METAG_ALPGGBFO |
| BGEO_SAMN07136937_METAG_ALPGGBFO_CGC7 | BGEO_SAMN07136937_METAG_ALPGGBFO |
| BGEO_SAMN07136947_METAG_PEBNHIGB_CGC5 | BGEO_SAMN07136947_METAG_PEBNHIGB |
| BGEO_SAMN07136947_METAG_PNJGMBLN_CGC11 | BGEO_SAMN07136947_METAG_PNJGMBLN |
| BGEO_SAMN07136947_METAG_PNJGMBLN_CGC4 | BGEO_SAMN07136947_METAG_PNJGMBLN |
| BGEO_SAMN07136947_METAG_PNJGMBLN_CGC6 | BGEO_SAMN07136947_METAG_PNJGMBLN |
| BGEO_SAMN07136957_METAG_PDLAKDGF_CGC17 | BGEO_SAMN07136957_METAG_PDLAKDGF |
| BGEO_SAMN07136957_METAG_PDLAKDGF_CGC18 | BGEO_SAMN07136957_METAG_PDLAKDGF |
| BGEO_SAMN07136957_METAG_PDLAKDGF_CGC19 | BGEO_SAMN07136957_METAG_PDLAKDGF |
| BGEO_SAMN07136957_METAG_PDLAKDGF_CGC21 | BGEO_SAMN07136957_METAG_PDLAKDGF |
| BGEO_SAMN07136957_METAG_PDLAKDGF_CGC22 | BGEO_SAMN07136957_METAG_PDLAKDGF |
| BGEO_SAMN07136958_METAG_IPBGLHHA_CGC5 | BGEO_SAMN07136958_METAG_IPBGLHHA |
| BGEO_SAMN07136960_METAG_CGNNLGDJ_CGC3 | BGEO_SAMN07136960_METAG_CGNNLGDJ |
| GCA_000009785.1_ASM978v1_genomic_CGC14 | GCA_000009785.1_ASM978v1_genomic |
| GCA_000009785.1_ASM978v1_genomic_CGC16 | GCA_000009785.1_ASM978v1_genomic |
| GCA_000009785.1_ASM978v1_genomic_CGC17 | GCA_000009785.1_ASM978v1_genomic |
| GCA_000009785.1_ASM978v1_genomic_CGC20 | GCA_000009785.1_ASM978v1_genomic |
| GCA_000009785.1_ASM978v1_genomic_CGC9 | GCA_000009785.1_ASM978v1_genomic |
| GCA_000012805.1_ASM1280v1_genomic_CGC12 | GCA_000012805.1_ASM1280v1_genomic |
| GCA_000012805.1_ASM1280v1_genomic_CGC15 | GCA_000012805.1_ASM1280v1_genomic |
| GCA_000023865.1_ASM2386v1_genomic_CGC22 | GCA_000023865.1_ASM2386v1_genomic |
| GCA_000023865.1_ASM2386v1_genomic_CGC24 | GCA_000023865.1_ASM2386v1_genomic |
| GCA_000023865.1_ASM2386v1_genomic_CGC26 | GCA_000023865.1_ASM2386v1_genomic |
| GCA_000023865.1_ASM2386v1_genomic_CGC30 | GCA_000023865.1_ASM2386v1_genomic |
| GCA_000025905.1_ASM2590v1_genomic_CGC10 | GCA_000025905.1_ASM2590v1_genomic |
| GCA_000025905.1_ASM2590v1_genomic_CGC12 | GCA_000025905.1_ASM2590v1_genomic |
| GCA_000025905.1_ASM2590v1_genomic_CGC13 | GCA_000025905.1_ASM2590v1_genomic |
| GCA_000025905.1_ASM2590v1_genomic_CGC14 | GCA_000025905.1_ASM2590v1_genomic |
| GCA_000025905.1_ASM2590v1_genomic_CGC2 | GCA_000025905.1_ASM2590v1_genomic |
| GCA_000025905.1_ASM2590v1_genomic_CGC27 | GCA_000025905.1_ASM2590v1_genomic |
| GCA_000025905.1_ASM2590v1_genomic_CGC29 | GCA_000025905.1_ASM2590v1_genomic |
| GCA_000025905.1_ASM2590v1_genomic_CGC4 | GCA_000025905.1_ASM2590v1_genomic |
| GCA_000025905.1_ASM2590v1_genomic_CGC5 | GCA_000025905.1_ASM2590v1_genomic |
| GCA_000025905.1_ASM2590v1_genomic_CGC9 | GCA_000025905.1_ASM2590v1_genomic |
| GCA_000092205.1_ASM9220v1_genomic_CGC15 | GCA_000092205.1_ASM9220v1_genomic |
| GCA_000092205.1_ASM9220v1_genomic_CGC20 | GCA_000092205.1_ASM9220v1_genomic |
| GCA_000092205.1_ASM9220v1_genomic_CGC5 | GCA_000092205.1_ASM9220v1_genomic |
| GCA_000144645.1_ASM14464v1_genomic_CGC5 | GCA_000144645.1_ASM14464v1_genomic |
| GCA_000147355.1_ASM14735v1_genomic_CGC3 | GCA_000147355.1_ASM14735v1_genomic |
| GCA_000147355.1_ASM14735v1_genomic_CGC4 | GCA_000147355.1_ASM14735v1_genomic |
| GCA_000147355.1_ASM14735v1_genomic_CGC7 | GCA_000147355.1_ASM14735v1_genomic |
| GCA_000148425.1_ASM14842v1_genomic_CGC1 | GCA_000148425.1_ASM14842v1_genomic |
| GCA_000148425.1_ASM14842v1_genomic_CGC13 | GCA_000148425.1_ASM14842v1_genomic |
| GCA_000148425.1_ASM14842v1_genomic_CGC4 | GCA_000148425.1_ASM14842v1_genomic |